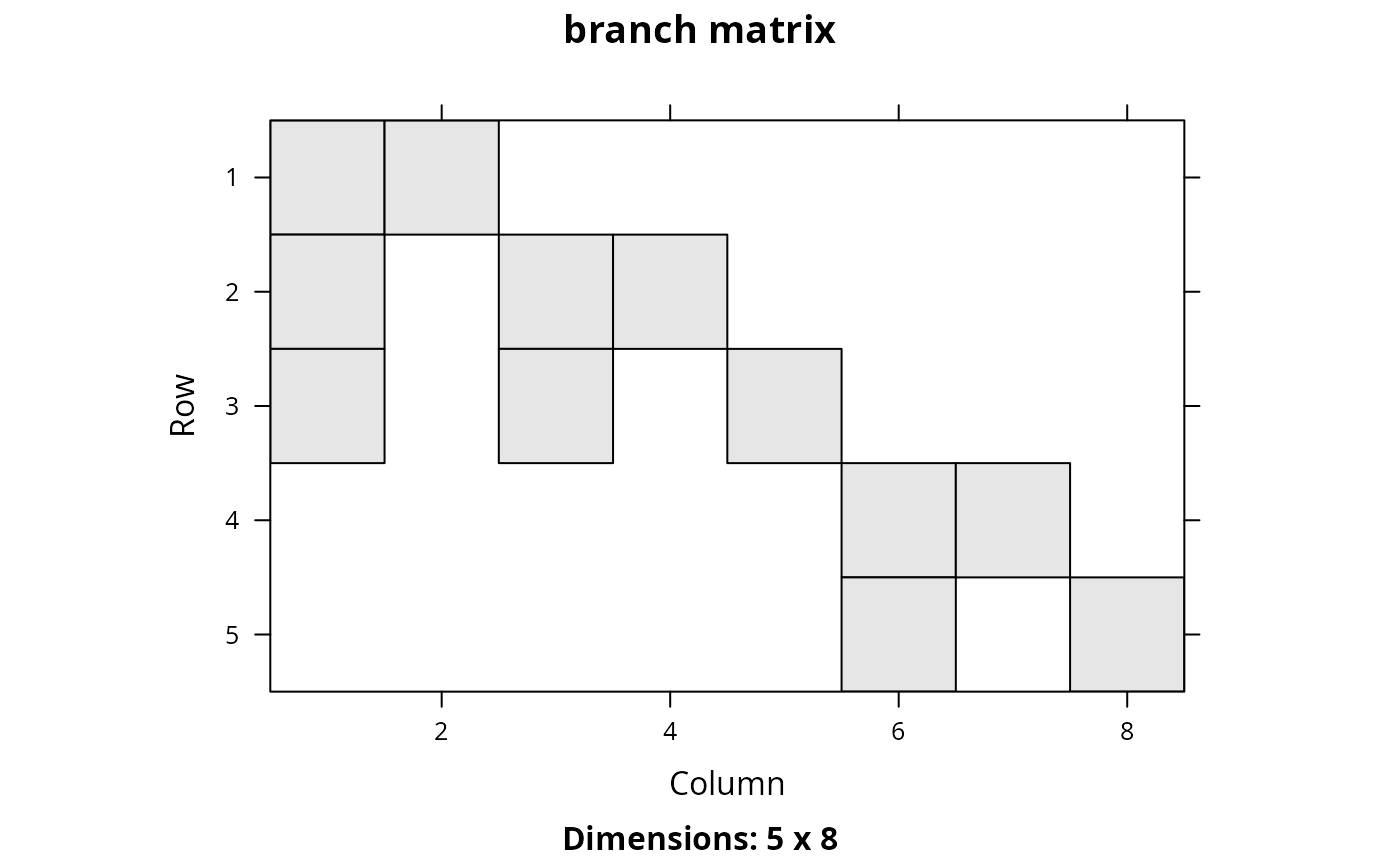
Phylogenetic trees depict the evolutionary relationships between different species. Each branch in a phylogenetic tree represents a period of evolutionary history. Species that are connected to the same branch both share that same period of evolutionary history. This function creates a matrix that shows which species are connected with branch. In other words, it creates a matrix that shows which periods of evolutionary history each species have experienced.
Usage
branch_matrix(x)
# Default S3 method
branch_matrix(x)
# S3 method for class 'phylo'
branch_matrix(x)Arguments
- x
ape::phylo()tree object.
Value
A Matrix::dgCMatrix sparse matrix object. Each row
corresponds to a different species. Each column corresponds to a different
branch. Species that inherit from a given branch are denoted with a one.
Examples
# load data
sim_phylogeny <- get_sim_phylogeny()
# generate species by branch matrix
m <- branch_matrix(sim_phylogeny)
# plot data
plot(sim_phylogeny, main = "phylogeny")
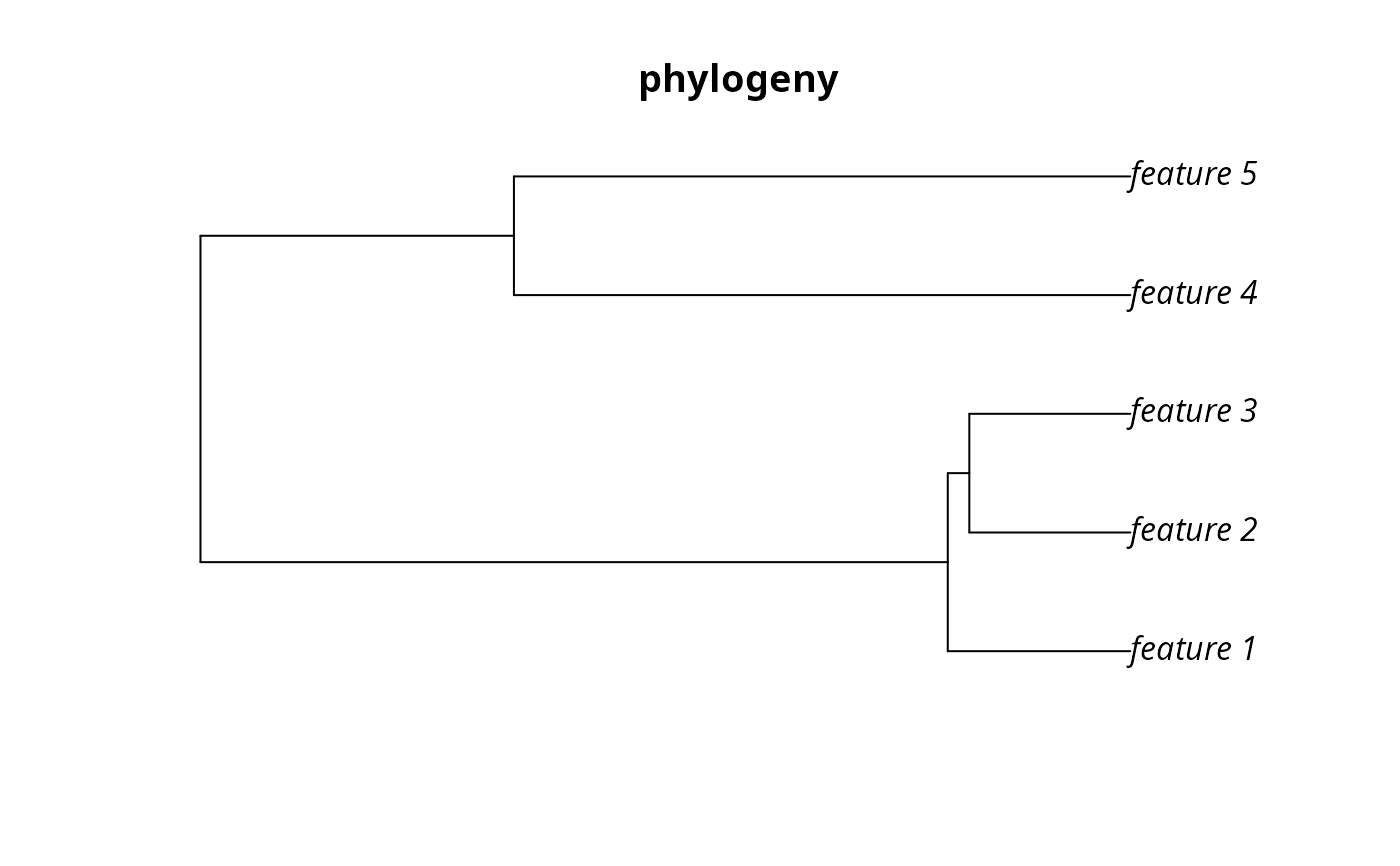 Matrix::image(m, main = "branch matrix")
Matrix::image(m, main = "branch matrix")